MGIS content
We are continuing to enrich the content of MGIS as well as adding new features. We have now 29 collections providing data to MGIS for a total of 6424 accessions.
The latest release includes additional Passport Data from new partners who are signatory of the MGIS Data sharing Agreement (DSA). We would like to thank for their confidence in providing their accession information the following:
- Cuba through the Instituto de Investigaciones de Viandas Tropicales (INIVIT),
- Philippines through Institute of Plant Breeding - University of the Philippines Los Baños (IPB-UPLB) with two collection sites Pili Drive 1 and Pili Drive 2,
- Sudan through the Agricultural Plant Genetic Resources Conservation and Research Center (APGRC),
- Venezuela through the Instituto Nacional de Investigaciones Agrícolas - Centro Nacional de Investigaciones Agropecuarias (INIAP-CENIAP)
Moreover, we have updated the Passport Data of the Bioversity International Musa Germplasm Transit Centre (ITC) in Belgium. 20 new accessions have been added to the ITC collection. Among which 5 accessions have been collected during a 16-day mission in October 2016 on the islands of Bougainville, Buka and Sohano in the Autonomous Region of Bougainville, Papua New Guinea (PNG). The mission was co-organized by the National Agriculture Research Institute (NARI) in PNG and Bioversity International, and was funded by the Genebanks CGIAR Research Programme, with a contribution of the Belgium government through the PhenSeeData project. You can look at the catalogue here.
Map
Since last year, after the changes related to the term of uses of google map services we switched all our maps to openstreet map.
Publications
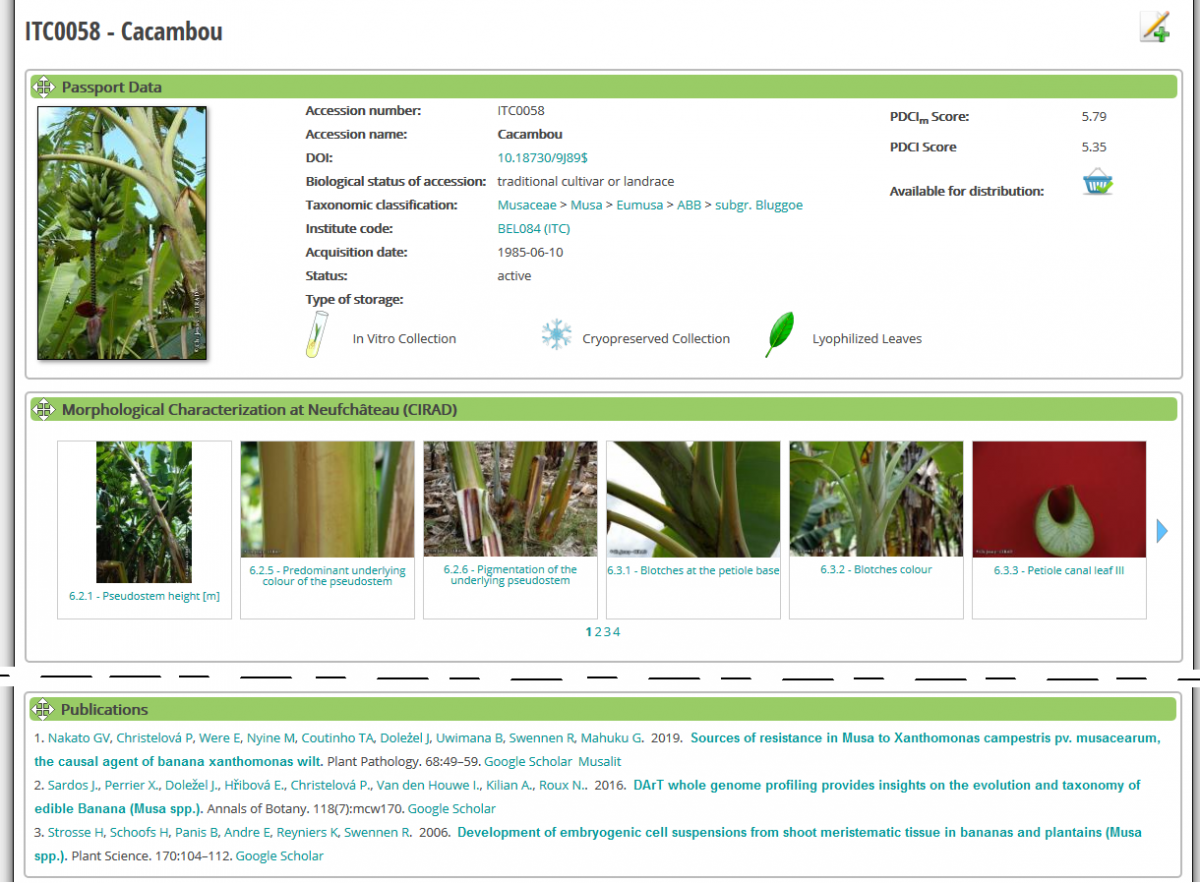
For two years, on a regular basis, scientific articles citing ITC accessions have been curated and linked to indiviudal accession pages. On each bottom of the page of ITC accessions you will find the list of publications citing this specific accession (see below).
Look to In-situ observations
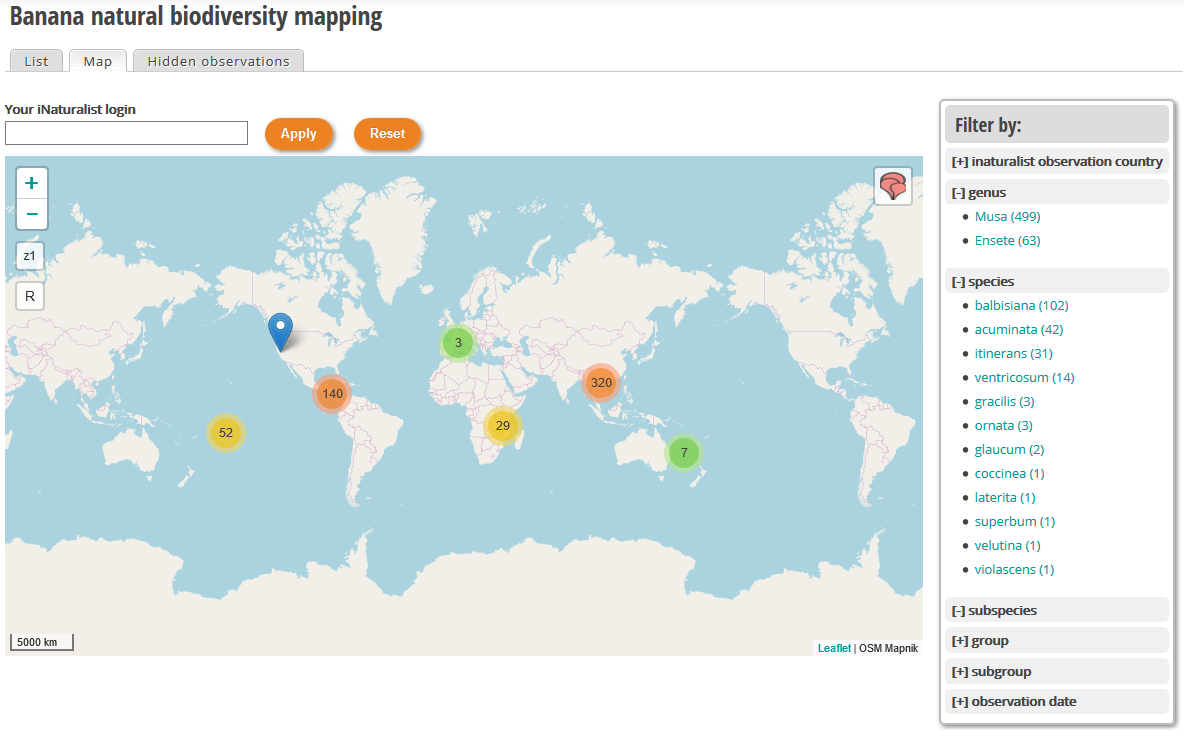
Since the end of the last year, we have worked on integrating data collected in the framework of the crowd sourcing project called "Banana natural diversity mapping" hosted on iNaturalist web site managed by Christophe Jenny (CIRAD). The objective of this module is to search and map observations and in a second phase to use this dataset to get a better overview of Musa spp. distribution at last crossing ex-situ and in-situ data could help to identify gaps in ex-situ collection. You can access to the module here.
DOI
MGIS continue to promote the use of DOI in collection. Since 2018, Bioversity international Musa Germplasm Transit Centre (ITC) assigns on routine basis DOI to material introduced in the collection.
DOIs are used as Permanent Unique Identifiers (PUID) in the context of the Global Information System (GLIS) of Article 17 of the International Treaty on Plant Genetic Resources for Food and Agriculture (ITPGRFA).
The SMTA generated once you complete an online request includes now the DOI next to the ITC accession code. This is to encourage researchers and other users to use the DOI in any publication they plan to release.
We have a dedicated page on DOI, accessible from the ABOUT menu or from the logo  visible at the bottom left side of the pages.
visible at the bottom left side of the pages.
Gigwa
The Gigwa module used to manage SNP datasets have been upgraded with the version 2 (Sempere et al, 2019). This new version of Gigwa provides a more intuitive and more powerful way to explore large amounts of genotyping data by offering a scalable solution to search for genotype patterns, functional annotations, or more complex filtering. Furthermore, its user-friendliness and interoperability make it widely accessible to the life science community. https://www.crop-diversity.org/mgis/gigwa
Breeding API.
The Breeding API (BrAPI) have also been updated to be compatible with version 1.3. It allows other information system to facilitate query and integration of MGIS data into their web site. You can query it online by going to this page: https://www.crop-diversity.org/mgis/brapi/query and an overview of the API is accessible here https://www.crop-diversity.org/mgis/brapi/overview.
As usual, comments and remarks are welcome.
Happy browsing, the MGIS Team.
